Research Overview⌗
In the Walker group, we are interested in how to connect biological phenomena and measurements of fluorescence using computational chemistry. Molecular modeling provides ways of translating between what we see in experiment, usually a signal or set of signals, to atomic or electronic level details of the system. To investigate fluorescence in biological systems, we use and develop new techniques in quantum chemistry, molecular dynamics, and excited state simulations. This has direct applications in imaging, both in medicine and in basic science.
Fluorescent Nucleotides and Aptamers⌗
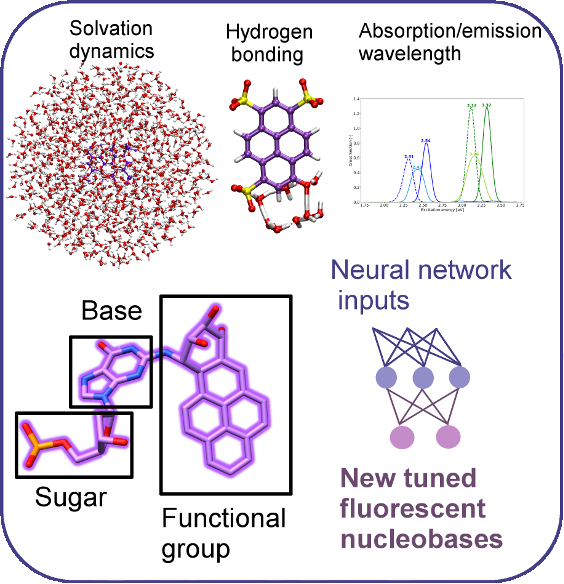
Theoretical chemistry provides avenues of testing large numbers of compounds and hypotheses in a systematic and inexpensive way, and also allows us to uncover principles for further design. One direction the Walker group will take is a bottom-up approach to designing new fluorescent biomolecules that absorb and emit brightly in the red part of the spectrum in order to image deep tissue. We will investigate a set of fluorescent synthetic nucleobases using quantum chemistry and molecular dynamics, and create a large data set that can be used for machine learning and design of new bases with desired properties, including shifting wavelengths to be more red and fluorescence quantum yield to be higher. These will then be tested in series, working up to large aptamer drugs like IL6SOMAmer, as pictured, in solution and bound to protein targets. Efficiently performing these calculations and interpreting such a dataset will require the development of new methodologies and approaches, particularly since fluorescence occurs on the nanosecond timescale.
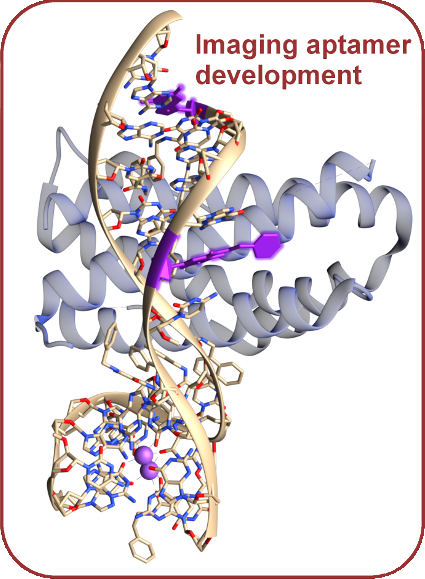
Aptamers, or short sequences of nucleotides, are easy to synthesize and are highly specific to their targets. By mutating specific parts of existing tight-binding aptamer drugs, we can potentially predict functionalization locations and effects of adding fluorescent synthetic nucleotides to various points in existing aptamer drugs, allowing for potential control and improved interpretation of experimental signals. Interleukin-6 is a protein implicated in cancer metastasis and inflammation, and provides an initial test system. We are also interested in Mango, an existing fluorescent aptamer that binds to biotin.
Reversible Photoswitchable Protein Design⌗
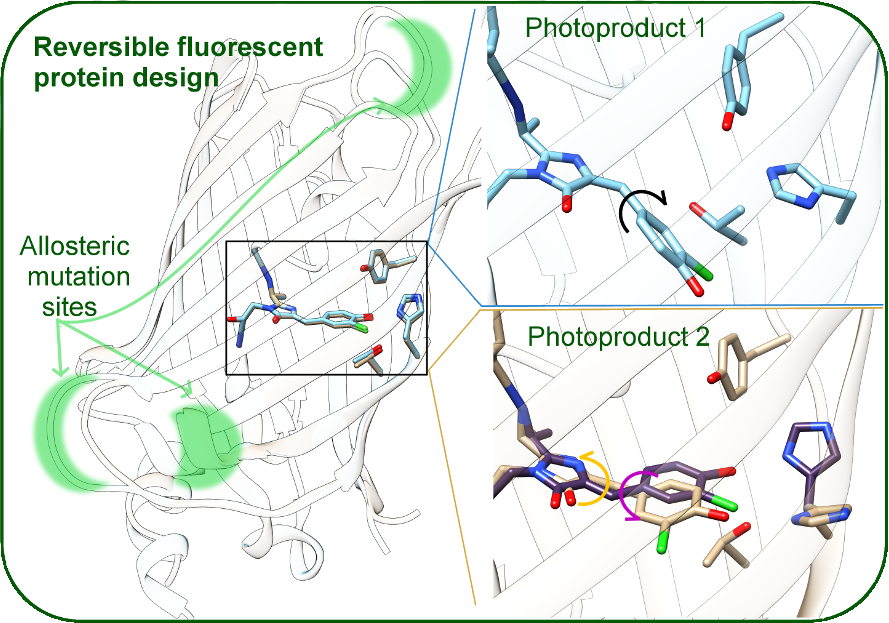
Fluorescent proteins and their derivatives are powerful experimental tools. They can be used directly as fluorescent tags, but also have applications as reversible photoswitches, where the focus is on nonradiative decay pathways that result in a physical change upon excitation with light at various wavelengths. Our group applies classical and combined quantum/molecular mechanics methods on the ground and excited state for the rational design of fluorescent proteins (FPs), based on a comprehensive theoretical picture of the relationship between excited state chromophore behavior, protein structure, and environment. Fluorescent proteins are ubiquitous as sensors in biochemical imaging, but the relationship between protein structures and motion, environmental factors such as pH, chromophore excited state behavior, and the actual measured fluorescent signal is still not fully understood. Since the connection between targeted mutations and experimental spectroscopic outcomes is not predictable, designing new FP-based sensors is an ongoing challenge. We are especially interested in specific challenges associated with creating bright, red, monomeric or photodissociative sensors for specific anions of interest, combining theoretical methodology and experimental collaborations where appropriate. Our work will also give further insight into the connection between atomic and electronic-level details of protein structure and measured spectroscopic signals. Supported by NSF CHE2338804.
FRET Tag Simulations of RNA Pol II⌗
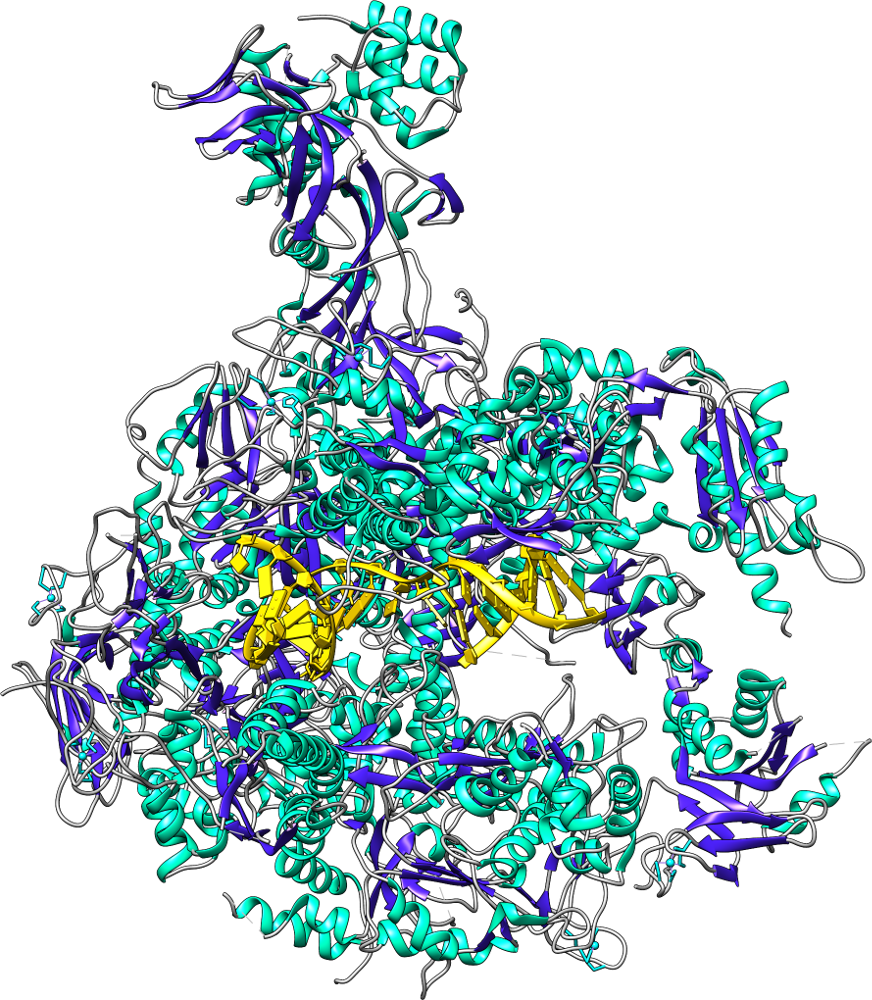
Another direction will be to investigate the effects of protein environment on fluorescence. Many experiments use fluorescent tags to investigate protein dynamics. This direction of the Walker group will involve investigating large protein systems with their fluorescent tags attached, both to compare fluorescent emission wavelengths and brightness to experimental quantities and to potentially design custom tags with new properties. As the tags become close to each other, energy transfers from Tag 1 to Tag 2, resulting in an immediate shift in emission wavelength. This can be used to compare the motions of proteins simulated on the computer with the motions obtained in experiment, while also providing greater detail on what is happening on the atomic level for even very large systems.